18.15: Chapter 15- A Gene Essential for Speech
- Page ID
- 113232
This page is a draft and under active development. Please forward any questions, comments, and/or feedback to the ASCCC OERI (oeri@asccc.org).
- Explain the first evidence that the FOXP2 gene was involved in human speech
- Describe evidence that FOXP2 is involved in vocalization in other species
- Describe the single nucleotide mutation on the FOXP2 gene, and the resulting alteration of the FOXP2 protein, which causes language difficulties
- Describe the general location of the FOXP2 gene in humans by identifying the chromosome where it is found
- Identify the differences between the human FOXP2 gene and the same gene in representative mammals and non-human primates
Overview
FOXP2 was the first gene discovered to be essential for human speech. It is expressed in many areas of the brain, including the basal ganglia and inferior frontal cortex, where it is essential for brain maturation and speech and language development (Enard et al., 2002). FOXP2 is found in many vertebrates ranging mouse to alligator and is involved in the vocalizations of many animals. For example, it is found in song birds and is important in their production of birdsong. In bats, FOXP2 is involved in echolocation. The gene has been highly conserved in mammals and few differences in the gene exist among mammal species. The protein that the gene produces is nearly identical in mice and primates.
The FOXP2 Gene
The FOXP2 (short for “forkhead box P2”) gene is the first gene that scientists ever associated with the human ability to speak (Nudel & Newbury, 2013). FOXP2 was first discovered in the KE family (medical name for this British family), about half of whose members had specific language impairments. Towards the end of 1980's, seven children of the family attended a special educational needs unit at a primary school in London. The head of the special needs unit discovered that the family had a speech disorder for three generations. Of the 30 family members, about half suffer from severe language deficiency, some are affected mildly, and few are unaffected (Watkins, et al., 1999). Their faces show rigidity at the lower half, and most cannot complete pronouncing a word. Many of them have severe stuttering and limited vocabulary. In particular, they have difficulty with consonants, and omit them, such as "boon" for "spoon", "able" for "table", and "bu" for "blue." Linguistic deficiency is also noted in written language both in reading and writing. They are characterized by lower nonverbal IQ in addition to their language difficulties. The first scientific report on the family's disorder, by Hurst, et al. (1990), showed that 16 family members were affected by severe abnormality, though their hearing was normal and some had normal intelligence, and that the condition was genetically inherited and was autosomal dominant.
Figure \(\PageIndex{1}\): FOXP2 protein and DNA. Forkhead box protein P2 (FOXP2) is a protein that, in humans, is encoded by the FOXP2 gene. FOXP2 is a member of the forkhead box family of transcription factors, proteins that regulate gene expression by binding to DNA. It has multiple effects and is expressed in the brain, heart, lungs and digestive system. Ribbon diagram of forkhead box P2 (FOXP2) protein. (Image from Wikimedia Commons; File:FOXP2 (2as5).png; https://commons.wikimedia.org/wiki/F...XP2_(2as5).png; by SWISS-MODEL, based on 2as5 from PDB; licensed under the Creative Commons Attribution-Share Alike 4.0 International license).
Using positron emission tomography (PET) and magnetic resonance imaging (MRI), Vargha-Khadem, et al. (1998) found that some brain regions were underactive (compared to baseline levels) in the KE family members and that some were overactive, when compared to normal people. The underactive regions included motor neurons that control face and mouth regions. The areas that were overactive included Broca's area, the speech center (which might indicate that the speech center is having to work much harder than in normal people in order to produce even faulty speech).
Fisher, et al. (1998) identified the exact location of the gene on the long arm of chromosome 7 (7q31). The chromosomal region (locus) was named SPCH1 (for speech-and-language-disorder-1), and it contains 70 genes. Using the known gene location of speech disorder from a boy, designated CS, of an unrelated family, they discovered in 2001 that the main gene responsible for speech impairment in both KE family and CS was FOXP2, and that this gene plays a major role in the origin and development of language (Lai, et al., 2001). Mutations in the gene result in speech and language problems (Vargha-Khadem & Liegeois, 2007) as seen in the KE family.
The exact problems caused by mutations in this gene remain hard to identify. This is not surprising when you consider the family of genes to which this one belongs. The FOX family of genes are transcription factors, which means that they produce proteins that can regulate the expression of a number of other genes by binding directly to their DNA. (The binding ability of these particular proteins comes from their forked shape, from which the gene family gets its name.) The FOXP2 gene would appear to play an important role in orchestrating the establishment of the neural pathways during embryonic development, some of which are required for normal vocalizations in several species studied, including humans. Surprisingly, this gene is extremely well preserved phylogenetically (across species): the protein that it produces is almost identical in mice and in primates, which are separated by some 130 million years of evolution.
The protein that the FOXP2 gene produces in humans differs by only two or three amino acids from the protein that it produces in other species. It is very likely these two or three amino acids make the difference between animals that cannot speak and humans who can. Moreover, the mutations that caused this difference are estimated to have occurred between 100,000 and 200,000 years ago, roughly the time that articulate language may have first emerged in human beings.
By performing a detailed analysis of the defective FOXP2 gene sequence in several members of the KE family, scientists were also able to identify the precise site of the mutation that caused this gene to malfunction in these individuals. This mutation occurs on exon 14 of the FOXP2 gene, when the guanine in a nucleotide is replaced by an adenine. As it happens, the part of the gene where this mutation occurs is precisely the one that codes for the “forkhead” portion of the protein—the part that binds to the DNA on other genes. This change in a single nucleotide on the FOXP2 gene has a direct impact on this protein, causing the amino acid arginine to be replaced with a histidine.
In the hundreds of normal subjects tested, the protein produced by FOXP2 always has an arginine at this particular site, while in the members of the KE family who suffered from specific language impairments, it always had a histidine. Hence there is not a shadow of a doubt about the mutation that causes this disorder. That said, it is still amazing to think that the mutation of a single one of the 2,500 nucleic bases in the FOXP2 gene is sufficient to impair language!
FOXP2 is also required for proper lung development, not only in humans but in other species as well. Additional research showed that Knockout mice with only one functional copy of the FOXP2 gene have significantly reduced vocalizations as pups (Shu et al., 2005). Knockout mice with no functional copies of FOXP2 are runted, display abnormalities in brain regions such as the Purkinje layer, and die an average of 21 days after birth from inadequate lung development.
Figure \(\PageIndex{2}\): The FOXP2 gene is located on the long (q) arm of chromosome 7 at position 31. More precisely, the FOXP2 gene is located from base pair 114,086,309 to base pair 114,693,771 on chromosome 7 (Image and caption from Wikimedia Commons; File:FOXP2 location.png; https://commons.wikimedia.org/wiki/F...2_location.png; by U.S. National Library of Medicine; in the public domain).
Evolution
The FOXP2 Gene is Found in Many Diverse Species
The FOXP2 gene is highly conserved in mammals (Enard et al., 2002). Conserved, in the evolutionary context, means that a trait or gene and its variants are found in a range of diverse species over long periods of evolutionary time suggesting that once it appeared it was retained as new species diverged from original ancestral species. The human gene differs from that in non-human primates by the substitution of just two amino acids, a threonine to asparagine substitution at position 303 (T303N) and an asparagine to serine substitution at position 325 (N325S) (Preuss, 2012). In mice it differs from that of humans by three substitutions, and in zebra finch by seven amino acids (Enard et al., 2002; Haesler et al., 2004; Teramitsu et al., 2004). One of the two amino acid differences between human and chimps also arose independently in carnivores and bats (Li et al., 2007; Shu et al., 2007). Similar FOXP2 proteins can be found in songbirds, fish, and reptiles such as alligators (Webb et al., 2005; Scharff & Haesler, 2005).
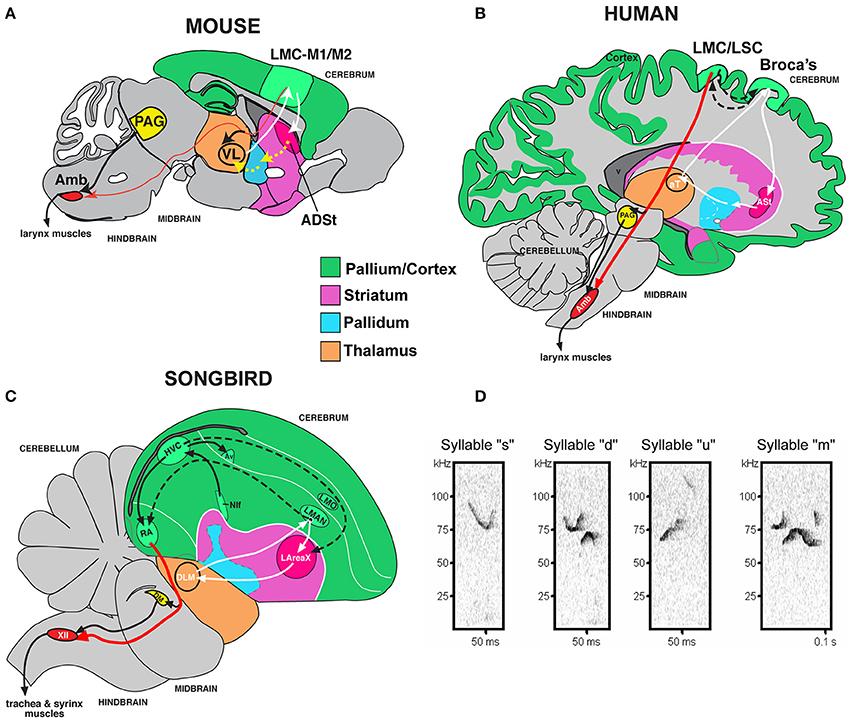
Figure \(\PageIndex{3}\): Mouse song system anatomy and syllable types. (A) Proposed anatomy of the rudimentary mouse forebrain vocal communication circuit based on Arriaga et al. (2012). Not shown are other connected brainstem regions, the amygdala, and insula. (B) Comparison with human, based on Arriaga et al. (2012) and Pfenning et al. (2014). (C) Comparison with songbird. (D) Sonograms of examples syllables of the four syllable categories quantified from a C57 male mouse USV song, labeled according to pitch jumps. Anatomical abbreviations: ADSt, anterior dorsal striatum; Amb, nucleus ambiguous; ASt, anterior striatum; aT, anterior thalamus; Av, nucleus avalanche; HVC, a letter-based name; LArea X, lateral Area X; LMO, lateral mesopallium oval nucleus; LMAN, lateral magnocellular nucleus of the nidopallium; LMC, laryngeal motor cortex; LSC, laryngeal somatosensory cortex; M1, primary motor cortex; M2, secondary motor cortex; NIf, interfacial nucleus of the nidopallium; PAG, periaqueductal gray; RA, robust nucleus of the arcopallium; T, thalamus; VL, ventral lateral nucleus of the thalamus; XIIts, 12th vocal motor nucleus, tracheosyringeal part (Figure and caption from Chabout et al., 2016).
The FOXP2 Gene Found in Neanderthal Fossils
DNA sampling from Homo neanderthalensis bones indicates that their FOXP2 gene is a little different though largely similar to those of Homo sapiens (i.e. humans) (Krause et al., 2005; Zimmer, 2016). Previous genetic analysis had suggested that the H. sapiens FOXP2 gene became fixed in the population around 125,000 years ago (Benítez-Burraco et al., 2008). Some researchers consider the Neanderthal findings to indicate that the gene instead swept through the population over 260,000 years ago, before our most recent common ancestor with the Neanderthals (Benítez-Burraco et al., 2008). Other researchers offer alternative explanations for how the H. sapiens version would have appeared in Neanderthals living 43,000 years ago (Benítez-Burraco et al., 2008).
Positive Selection for FOXP2
According to a 2002 study (Enard, et al., 2002), the FOXP2 gene showed indications of recent positive selection (Enard, et al., 2002; Toda et al., 1992). Some researchers have speculated that positive selection is crucial for the evolution of language in humans (Enard, et al., 2002).
The figure below (Figure 14.15.4) shows three types of selection. The top shows positive selection. Positive selection is also known as Darwinian selection and is the type of selection that Darwin envisioned as the primary mechanism of evolution (for more on the types of selection, see Module 3.5). The original population distribution, before selection, is shown in red. The population distribution after selection is shown in blue. In directional evolution, the population values for the trait move toward one extreme of the trait. By positive selection, a new adaptive trait can sweep through a population. However, "though advantageous mutations are of great interest, they are difficult to detect and analyze because of the fact that neutral and deleterious mutations predominate them in frequency" (Thomas Lab, University of Washington).
Figure \(\PageIndex{4}\): These charts depict the different types of genetic selection. On each graph, the x-axis variable is the type of phenotypic trait and the y-axis variable is the number of organisms. Group A is the original population and Group B is the population after selection. Graph 1 shows directional selection, in which a single extreme phenotype is favored. Graph 2 depicts stabilizing selection, where the intermediate phenotype is favored over the extreme traits. Graph 3 shows disruptive selection, in which the extreme phenotypes are favored over the intermediate (Image and caption from Wikimedia Commons; File:Genetic Distribution.svg; https://commons.wikimedia.org/wiki/F...stribution.svg; by Ealbert17; licensed under the Creative Commons Attribution-Share Alike 4.0 International license).
Recent Positive Selection for FOXP2 Disputed
Others, however, were unable to find a clear association between species with learned vocalizations and similar mutations in FOXP2 (Webb et al., 2005; Scharff & Haesler, 2005). A 2018 analysis of a large sample of globally distributed genomes confirmed there was no evidence of positive selection, suggesting that the original signal of positive selection may be driven by sample composition (Atkinson et al., 2018; Williams, 2020). Insertion of both human mutations into mice, whose version of FOXP2 otherwise differs from the human and chimpanzee versions in only one additional base pair, causes changes in vocalizations as well as other behavioral changes, such as a reduction in exploratory tendencies, and a decrease in maze learning time. A reduction in dopamine levels and changes in the morphology of certain nerve cells are also observed (Enard et al., 2009).
Figure \(\PageIndex{5}\): Human FOXP2 gene and evolutionary conservation is shown in a multiple alignment (at bottom of figure) in this image from the UCSC Genome Browser. Note that conservation tends to cluster around coding regions (exons).
Migration of Human Groups and Genetic and Linguistic Relationships
Around the end of World War II, the Italian population geneticist Luca Cavalli-Sforza began constructing genealogical trees that established relationships among populations throughout the world. By cross-tabulating data on several dozen genes, Cavalli-Sforza established a relationship between American Indians and Asians. This finding is consistent with the most common theory about how the New World was populated: by peoples who crossed from Siberia to Alaska when the Bering Strait was frozen over during the last great Ice Age, some 30,000 years ago.
Cavalli-Sforza’s findings assumed even greater importance when he correlated them with analogous studies on languages. When Cavalli-Sforza compared the genealogical trees established by geneticists with those established by linguists, with just a few exceptions, he found that the people who speak each of the 15 major families of languages are genetically related as well. The explanation for this remarkable concordance is that when a population migrates to a new territory, it takes its genes along as well as its language.
But there have been many criticisms of Cavalli-Sforza’s approach, and in particular his way of defining a population. In the works of Cavalli-Sforza and his followers, the first step is to define a population, by linguistic criteria among others. Correlations are then established between these populations and their languages, which seems like a dangerously circular approach. It has also been noted that these studies are more convincing on a small scale or a large scale, but far less so on an intermediate one. The reason is that it is easier to distinguish Inuit from Bantu, for example, than to differentiate the various populations that speak Bantu languages.
In addition, the DNA samples used in many studies come from blood banks, and the accompanying records may be biased or false, because for various reasons, when people give blood, they may report their ethnicity as different from what it actually is. This points to the fact that errors in interpretation may be found when controls are applied to both linguistic and genetic data.
Summary
The FOXP2 gene is found widely in the animal kingdom and the human version of the gene differs little from that in other species, yet the human variant appears to have made language possible. Mutation of a single one of the 2,500 nucleic bases in the FOXP2 gene is sufficient to impair language. Though many researchers believe that the FOXP2 gene played an important role in the evolution of human language, evidence for positive selection for it is still controversial.
References
Atkinson EG, Audesse AJ, Palacios JA, Bobo DM, Webb AE, Ramachandran S, Henn BM (September 2018). "No Evidence for Recent Selection at FOXP2 among Diverse Human Populations". Cell. 174 (6): 1424–1435.e15
Benítez-Burraco A, Longa VM, Lorenzo G, Uriagereka J (November 2008). "Also sprach Neanderthalis... Or Did She?". Biolinguistics. 2 (2): 225–232.
Cavalli-Sforza, L. L. & Edwards, A. W. F. (1963) in Genetics Today, Proceedings of the 11th International Congress of Genetics, The Hague, The Netherlands, ed. Geerts, S. J. (Pergamon, New York), No. 3, pp. 923-933.
Cavalli-Sforza, L. L. (1986) in Evolutionary Perspectives and the New Genetics, eds. Gershowitz, H., Rucknagel, D. L. & Tashian, R. E. (Liss, New York), pp. 13-30.
Cavalli-Sforza, L. L., Piazza, A., Menozzi, P., & Mountain, J. (1988). Reconstruction of human evolution: bringing together genetic, archaeological, and linguistic data. Proceedings of the National Academy of Sciences, 85 (16), 6002-6006.
Chabout, J., Sarkar, A., Patel, S. R., Radden, T., Dunson, D. B., Fisher, S. E., & Jarvis, E. D. (2016). A Foxp2 mutation implicated in human speech deficits alters sequencing of ultrasonic vocalizations in adult male mice. Frontiers in behavioral neuroscience, 10, 197.
Enard W, Przeworski M, Fisher SE, Lai CS, Wiebe V, Kitano T, et al. (August 2002). "Molecular evolution of FOXP2, a gene involved in speech and language" (PDF). Nature. 418 (6900): 869–72.
Enard W, Gehre S, Hammerschmidt K, Hölter SM, Blass T, Somel M, et al. (May 2009). "A humanized version of Foxp2 affects cortico-basal ganglia circuits in mice". Cell. 137 (5): 961–71
Fisher, S. E.; Vargha-Khadem, F.; Watkins, K. E.; Monaco, A.P.; Pembrey, M. E. (1998). "Localisation of a gene implicated in a severe speech and language disorder". Nature Genetics. 18 (2): 168–170. doi:10.1038/ng0298-168
Haesler S, Wada K, Nshdejan A, Morrisey EE, Lints T, Jarvis ED, Scharff C (March 2004). "FoxP2 expression in avian vocal learners and non-learners". The Journal of Neuroscience. 24 (13): 3164–75.
Hurst, J. A.; Baraitser, M.; Auger, E.; Graham, F.; Norell, S. (1990). "An extended family with a dominantly inherited speech disorder". Developmental Medicine & Child Neurology. 32 (4): 352–355. doi:10.1111/j.1469-8749.1990.tb16948.x
Krause J, Lalueza-Fox C, Orlando L, Enard W, Green RE, Burbano HA, et al. (November 2007). "The derived FOXP2 variant of modern humans was shared with Neandertals". Current Biology. 17 (21): 1908–12.
Lai, C. S. L.; Fisher, S. E.; Hurst, J. A.; Vargha-Khadem, F.; Monaco, A. P. (2001). "A forkhead-domain gene is mutated in a severe speech and language disorder". Nature. 413 (6855): 519–523
Li G, Wang J, Rossiter SJ, Jones G, Zhang S (September 2007). Ellegren H (ed.). "Accelerated FoxP2 evolution in echolocating bats". PLOS ONE. 2 (9): e900.
Nudel, R., & Newbury, D. F. (2013). Foxp2. Wiley Interdisciplinary Reviews: Cognitive Science, 4 (5), 547-560.
Preuss TM (June 2012). "Human brain evolution: from gene discovery to phenotype discovery". Proceedings of the National Academy of Sciences of the United States of America. 109 (Suppl 1): 10709–16.
Scharff C, Haesler S (December 2005). "An evolutionary perspective on FoxP2: strictly for the birds?". Current Opinion in Neurobiology. 15 (6): 694–703
Shu, W., Cho, J. Y., Jiang, Y., Zhang, M., Weisz, D., Elder, G. A., ... & Buxbaum, J. D. (2005). Altered ultrasonic vocalization in mice with a disruption in the Foxp2 gene. Proceedings of the National Academy of Sciences, 102(27), 9643-9648.
Shu W, Lu MM, Zhang Y, Tucker PW, Zhou D, Morrisey EE (May 2007). "Foxp2 and Foxp1 cooperatively regulate lung and esophagus development". Development. 134 (10): 1991–2000.
Teramitsu I, Kudo LC, London SE, Geschwind DH, White SA (March 2004). "Parallel FoxP1 and FoxP2 expression in songbird and human brain predicts functional interaction". The Journal of Neuroscience. 24 (13): 3152–63.
Thomas, J.H., Thomas Lab. Genome Sciences, University of Washington; http://depts.washington.edu/jtlab/. Detection and analysis of positive selection. Retrieved 5/11/2022.
Toda M, Okubo S, Ikigai H, Suzuki T, Suzuki Y, Hara Y, Shimamura T (1992). "The protective activity of tea catechins against experimental infection by Vibrio cholerae O1". Microbiology and Immunology. 36 (9): 999–1001.
Vargha-Khadem, F.; Watkins, K. E.; Price, C. J.; Ashburner, J.; Alcock, K. J.; Connelly, A.; Frackowiak, R. S. J.; Friston, K. J.; Pembrey, M. E.; Mishkin, M.; Gadian, D. G.; Passingham, R. E. (1998). "Neural basis of an inherited speech and language disorder". Proceedings of the National Academy of Sciences. 95 (21): 12695–12700. doi:10.1073/pnas.95.21.12695. PMC 22893. PMID 9770548
Vargha-Khadem, F. & Liegeois, F. (2007). "From speech to gene: The KE family and the FOXP2". In Braten, Stein (ed.). On Being Moved : From Mirror Neurons to Empathy. Amsterdam: John Benjamins Publication. p. 11.
Watkins, Kate E; Gadian, David G.; Vargha-Khadem, Faraneh (1999). "Functional and Structural Brain Abnormalities Associated with a Genetic Disorder of Speech and Language". The American Journal of Human Genetics. 65 (5): 1215–1221. doi:10.1086/302631. PMC 1288272. PMID 10521285.
Webb, DM, Zhang J (2005). "FoxP2 in song-learning birds and vocal-learning mammals". The Journal of Heredity. 96 (3): 212–6.
Williams, S. (2020). "Language Gene Dethroned". The Scientist Magazine®. Retrieved 5/11/22.
Zimmer C (17 March 2016). "Humans Interbred With Hominins on Multiple Occasions, Study Finds". The New York Times. Retrieved 17 March 2016.
Attributions
Contributed by Kenneth A. Koenigshofer, PhD. adapted from Genes that are essential for speech by Bruno Dubuc under a Copyleft license.
Some text and images from Wikipedia, KE Family, and FOXP2, retrieved September 4, 2021. Text for section, Evolution, from Wikipedia, FOXP2.


